- RT theory allows us to explain the scattering & absorption of photons
- ... by describing the optical properties and structure of the scene
- However, we want to find out about the surface from the data!
- E.g. we want to infer LAI, chlorophyll, ... from reflectrance measurements
- The inverse problem....
The inverse problem¶
- An RT model $\mathcal{H}$ predicts directional reflectance factor, $\vec{\rho}_{m}(\Omega, \Omega')$
- $\dots$ as a function of a set of input parameters: LAI, chlorophyll concentration, equivalent water thickness...
- $\mathcal{H}$ combination leaf RT model, canopy RT model and soil RT model
- PROSPECT (Liberty?)
- SAIL (ACRM, Semidiscrete, ...)
- Linear mixture of spectra assuming Lambertian soil (Walthall, Hapke, ...)
- For this lecture, we'll refer to $\mathcal{H}$ as the observation operator
- Stack input parameters into vector $\vec{x}$.
- Other information e.g. illumination geometry, etc
- Our task is to infer $\vec{x}$ given observations $\vec{\rho}(\Omega, \Omega')$
- The model couples the observations and our parameters
- In some cases, we might be able to provide an analytic inversion
- However, we have ignored observational uncertainties
- We have also ignored the model uncertainty (inadequacy): a model is not reality
- These uncertainties will translate into uncertainty into our inference of $\vec{x}$
- Is there a framework for this?
- Least squares: minimise data-model mismatch
- Assuming observational noise,
- Any solution that falls within the error bars cannot be discarded
- Moreover, models very linear
- Haven't even considered model inadequacy!
- Eg simulate some healthy wheat canopy (LAI=4.2), and then run our RT model with random inputs
- Accept solutions that fall within the observational error... ($\Rightarrow$ brute force montecarlo!)
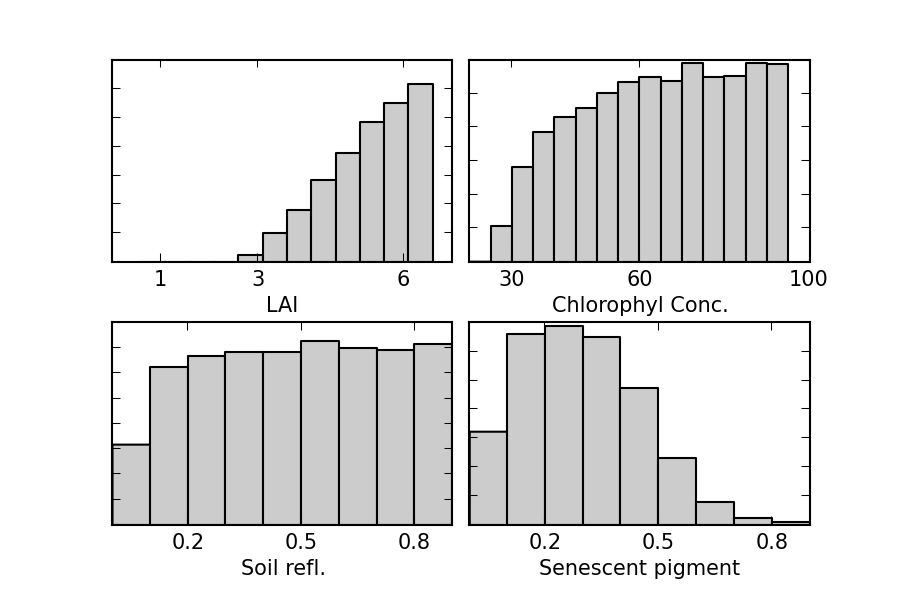
- Large uncertainty in estimate of state.
- $\Rightarrow$ limited information content in the data.
- Need more information..
- Add more observations
- Decrease observational uncertainty
- Need to consider sensitivity of the model.
Reverend Bayes to the rescue¶

- We assume that parameter uncertainty can be encoded if we treat $\vec{x}$ as a probability density function (pdf), $p(\vec{x})$.
- $p(\vec{x})$ encodes our belief in the value of $\vec{x}$
- Natual treatment of uncertainty
- We are interested in learning about $p(\vec{x})$ conditional on the observations $\vec{R}$, $p(\vec{x}|\vec{R})$.
- Bayes' Rule states how we can learn about $p(\vec{x}|\vec{R})$
- In essence, Bayes' rule is a statement on how to update our beliefs on $\vec{x}$ when new evidence crops up
$$
p(\vec{x} | \vec{R}, I ) =\frac{ p (\vec{R} | \vec{x}, I)\cdot p(\vec{x},I)}{p(\vec{R})}\propto p (\vec{R} | \vec{x}, I)\cdot p(\vec{x},I)
$$
- $p(\vec{R}|\vec{x},I)$ is the likelihood function
- encodes the probability of $\vec{R}$ given $\vec{x}$, and any other information ($I$)
- $p(\vec{x})$ is our a priori belief in the pdf of $\vec{x}$
- $p(\vec{R}$ can be thought of as normalisation constant, and we'll typically ignore it
- A way to picture Bayes' rule:
The prior $p(\vec{x})$¶
- Encodes everything we know about $\vec{x}$ before we even look at the data
- In some cases, we can have uninformative priors...
- ... but the real power is that it allows us to bring understanding, however weak to the problem!
The likelihood $p(\vec{R}|\vec{x})$¶
- The likelihood states is our data generative model
- It links the experimental results with the quantity of inference
- It includes our observations, their uncertainties, but also the model and its uncertainties
- Assume that our model is able perfect (ahem!), so if $\vec{x}$ is the true state, the model will predict the observation exactly
- Any disagreement has to be due to experimental error. We'll assume it's additive:
- Assume that $\epsilon \sim \mathcal{N}(\vec{\mu},\mathbf{\Sigma}_{obs})$
- For simplicity, $\vec{\mu}=\vec{0}$
The likelihood is then given by $$ p(\vec{R}|\vec{x})\propto\exp\left[-\frac{1}{2}\left(\vec{R}-\mathcal{H}(\vec{x})\right)^{\top}\mathbf{\Sigma}_{obs}^{-1}\left(\vec{R}-\mathcal{H}(\vec{x})\right)\right] $$
Have you ever seen this function?
- Maybe its univariate cousin...
- Can you say something about (i) the shape of the function and (ii) interesting points?
The posterior¶
- We can simply multiply the likelihood and prior to obtain the posterior
- In most practical applications with a non-linear $\mathcal{H}$, there is no closed solution
- However, if $\mathcal{H}$ is linear ($\mathbf{H}$), we can solve directly
- Simple 1D case, assuming Gaussian likelihood & prior:
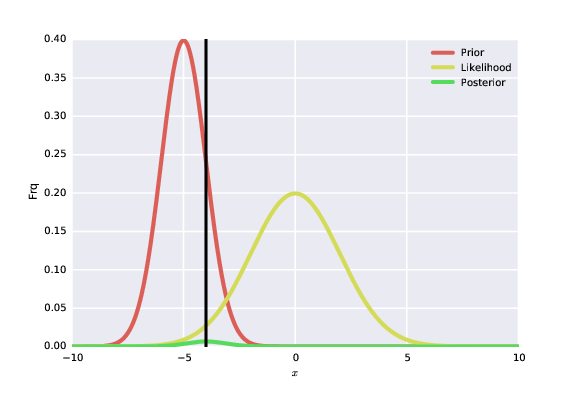
- What can you say about the information content of the data in this synthetic example?
Some simple 1D Gaussian maths...¶
- Try to infer $x$ from $y$, using an identity observation operator (i.e., $x=y$, so $\mathcal(x)=1$ and Gaussian noise:
- Assume that we only know that $x$ is Gaussian distributed with $\mu_p$ and $\sigma_0$:
- The posterior distribution is indeed a Gaussian, and its mean and std dev can be expressed as an update on the prior values
- Weighted by the relative weighting of the uncertainties
- If we now had a new measurement made available, we could use the posterior as the prior, and it would get updated!
Ill posed problems¶
- Stepping back from Bayes, consider the logarithm of the likelihood function:
- Maximum occurs when the model matches the observatios
- So we can just maximise the model-data mismatch and be done, right?
- Remember our generative model, and think what this implies:
- Formally, any solution that falls within the error bars is a reasonable solution
Priors¶
- Simplest prior constraints might encode
- range of parameters
- physical limits of parameters (e.g. a mass must be $>0$)
- More sophisticated priors might encode more subtle information such as
- expected values from an expert assessment
- expected values from a climatology
- It is usually hard to encode the prior information as a pdf
- We tend to use Gaussians as easy to parameterise
Variational solution¶
- If prior(s) & likelihood Gaussian...
- And if all terms are linear
- $\Rightarrow$ can solve for posterior by minimising $J(\vec{x})$:
- You can add more constraints (more observations, more prior constraints...)
- Uncertainty given by inverse of Hessian @ minimum of $J(\vec{x})$
- Matrix of second derivatives
- Also works OK for weakly non-linear systems!
More subtle priors¶
- What about the expectation of
- spatial correlation of the state
- temporal correlation of the state?
- Can encode expectation of smoothness in the prior
- What if we have a model of e.g. LAI evolution with e.g. thermal time?
- ... Or a full-blown vegetation/crop model?
The concept of a model¶
- Smoothness constraint is a simple vegetation model
- LAI today = LAI tomorrow
- This temporal evolution model is WRONG
- More complex mode likely to also be wrong $\Rightarrow$ uncertainty!
- Encode "closeness to model" $\mathcal{M}(\vec{x})$ constraint as a Gaussian pdf: $$ p(\vec{x}|\mathcal{M},I_m)\propto -\frac{1}{2}\exp\left[ - (\vec{x} - \mathcal{M}(I_m)^{\top}\mathbf{C}_{model}^{-1}(\vec{x} - \mathcal{M}(I_m)\right] $$
Simplest model¶
- Smoothness $$ x_{k+1} = x_{k} + \mathcal{N}(0,\sigma_{model}) $$
We can encode this as a matrix, if e.g. we stack $x$ over time in a vector...: $$ \begin{pmatrix} 1 &-1& 0 & 0 & \cdots\\ 0 &1 &-1 & 0 & \cdots \\ \vdots & \vdots & \vdots & \vdots & \cdots \\ \end{pmatrix}\vec{x} $$
It's a linear form!
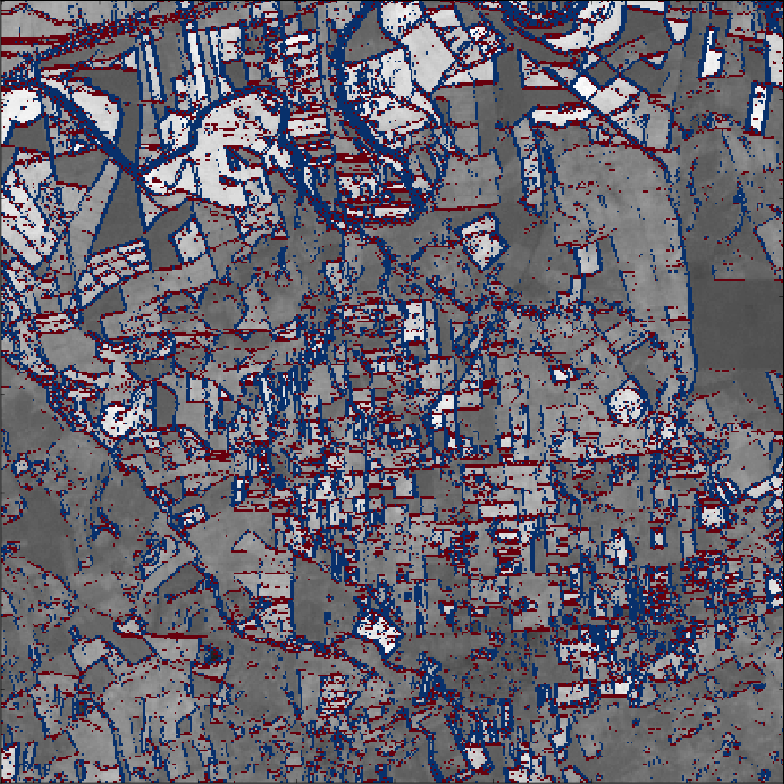
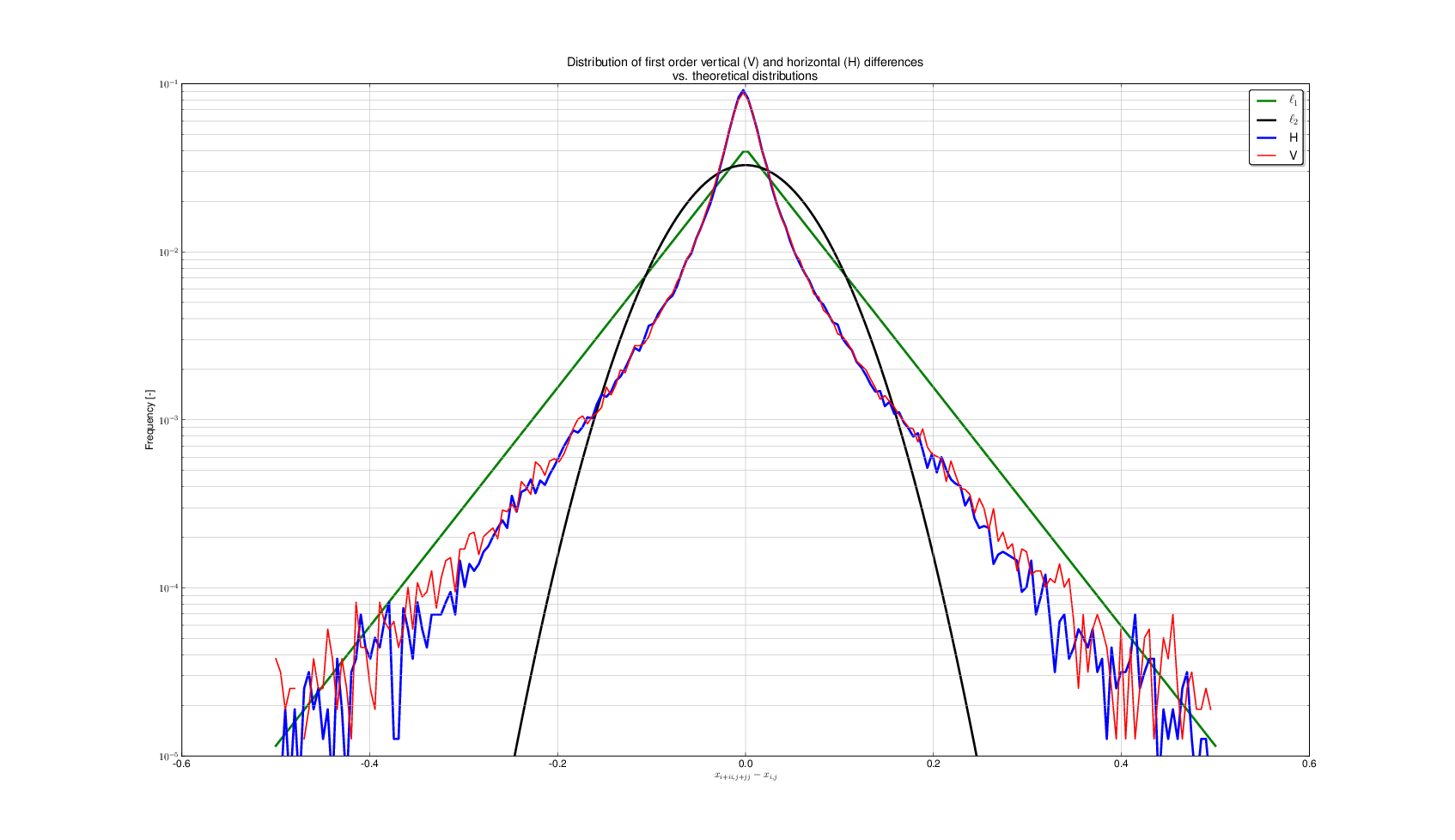
- Model can also be applied in space ($\Rightarrow$) Markov Random Field
- Vertical (red) and horizontal (blue) differences between neighbouring pixels
- Over a given threshold
eoldas_ng¶
- Python tool that allows you to build variational EO problems
- In development
- Main bottleneck: slow RT models
- Emulation of models
Final remarks¶
- Inversion of a model to infer input parameters is ill posed
- You need to add more information
- priors
- more observations
- Track all the uncertainties
- Physically consistent way of merging observations
- Variational solution can be efficient
- Apply smoothness and model constraints as priors
eoldas_ngtool
